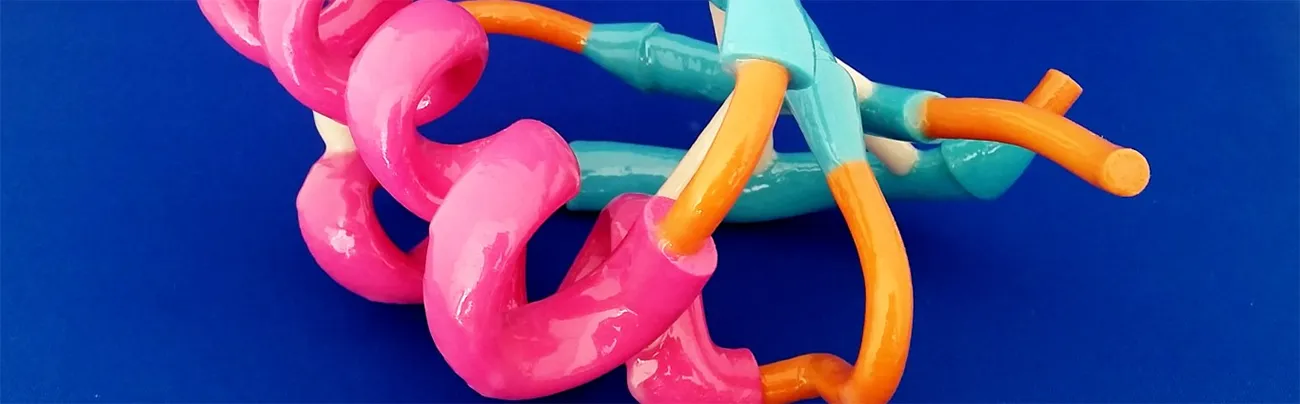

Over the last few decades, we have been able to significantly improve our understanding of the three-dimensional structure of large molecules, like nucleic acids and proteins, and how this relates to their function. With resources like the Protein Data Bank or the Nucleic Acid Database, in combination with modifiers such as Pymol, Chimera, JSmol and many more, 3D renders and animations of molecules can now be created in no time.
A recent trend is to not only view the digital models, but also 3D-print a physical version as well. In this article, we explain how to prepare your digital model for 3D printing.

Why 3D printed models of Proteins, Nucleic Acids, and other molecules?
While two-dimensional visualizations seem sufficient, there are a few use cases where it is more effective to have an actual physical model. For example, students – like most people – can struggle to imagine the three-dimensional structure based on two-dimensional images. Also, a physical model can be used to demonstrate the structure-function relationship of a molecule, e.g. ligand binding. Lastly, a physical model is an obvious focus point for presentations or exhibitions, allowing you to better communicate your findings.
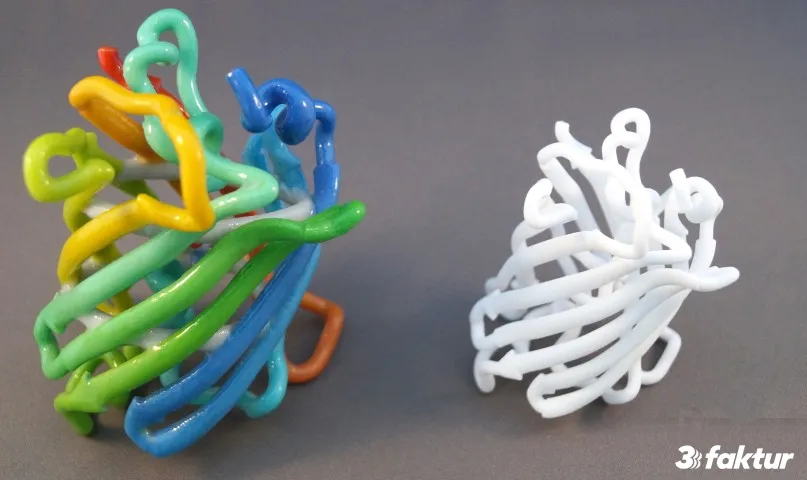

How to prepare a Protein 3D model for 3D printing
First you will need a digital model of your molecule of interest. If you’re lucky it might already be available in the expansive NIH 3D Printing exchange (As of this writing they have ~3,000). If your molecule is not part of the collection, you can follow the steps below to easily prepare your own (and make sure to post it later on NIH!).
There are multiple options to generate a 3D printable protein model. These instructions are written for the popular software Chimera, other resources are listed at the end of the page.
- Find your model on the Protein Data Bank and download in PDB Format (gz).
- In case you haven’t yet, download and install Chimera (Free for private use and academic or research institutes).
- Start Chimera,and open the file (File -> Open).
- Often, the model will contain additional chains or ligands. To remove the extra chains, do the following:
- Select the chain you do not need (Select -> Chain), go to Actions -> Ribbon -> Hide.
- If there are many extra chains, select the one you need (Select -> Chain), and then click on Select – Invert all models. This will select all chains besides the one you need. Then, go to Actions -> Ribbon -> Hide.
- For a ligand,
- Even if you want to print the ligand as well, we still recommend removing as described above and printing it separately. Since there is normally (logically) a gap between ligand and molecule they should ideally be printed as separate objects.
- Remove the ligand by selecting the entire chain (Select -> Chain), then go to Actions ->Atoms/Bonds -> Hide.
- Prepare the chain for 3D-printing: Most importantly, you need to make sure the structure is thick enough to survive printing. Optionally you can add colors to highlight features or communicate other important information.
- Thickness of the structures: In order to adjust the sizes, select the entire chain (Select -> Chain) and go to Tools -> Depiction -> Ribbon Style Editor. As a rule of thumb, the minimum thickness in smaller models should not be below 3 mm, in models with height, length or width greater than 20 cm the minimum thickness should be somewhere at least 6 mm, instructions on how to measure the wall thickness are at the end of this section.
- Colors (optional): color the secondary structures by doing the following: Select -> Structure -> Secondary Structure -> Coil. Go to Actions -> Color and chose a Coil color. Repeat the same process with Helices and Sheets.
- Done! Now export your model for 3D printing.
- Go to File -> Export Scene. For file format use STLfor non-colored 3D-printing (i.e for FDM, SLS, or SLA 3D printing technologies) and VRML (.wrl, .vrml) for colored models.
Most models are now ready to print. Ocassionally they may still have other issues. To check upload them to the free cloud-based software Netfabb.
Additional Information
How to measure the wall thickness with Meshmixer
Download and install Meshmixer (free). Load your molecule model. Click on analysis (left toolbar) -> measure – > Type and select the first item (it is already selected, it looks like a diameter). The chose in the very same menu Direction -> The icon with ‘n’ (already selected). Click on some parts of your model to get an impression of the wall thickness of the structure you’ve created. If it is too thin, go back to Chimera and increase the thickness.
How to extract your Ligand
Select the chain of interest, go to Actions ->Ribbon -> Hide. This removes the macro molecule. Now go to Actions -> Atoms/Bonds -> Show. If you haven’t so already, now you are now able to see the ligand. Now go to Actions -> Atoms/Bonds -> and choose the style, e.g., ball & stick or sphere (those are the most common types for 3D printing). You’re all set, you can extract the ligand as described below.
Print your 3D Protein Model
Once you have a printable protein model, it’s time to print it! In general, there are four technologies that are most commonly used for molecule printing:
- FDM: That’s the Ultimakers, Makerbots, Leapfrogs, Zortraxes and self-assembled 3D-printer kits that you can find at most Universities and academic Institutions nowadays. They basically melt a string of plastic to create a physical object layer by layer. Printing on these machines is cheap but the quality is rough. However, for teaching purposes it is often sufficient. Hanging sections of your model will require a support structure – this must be manually removed after printing – further compromising surface quality.
- SLA (Stereolithography): Just like FDM, this technology requires support structures, therefore it is not the best for protein models. But it is a great technology to print simple molecule structures. Compared to FDM, the surfaces are very smooth and the print is highly accurate. We at 3Faktur print most ligands with this technology. Some institutions may also have such printers at their disposal (e.g. Formlabs). If not, there are many 3D-printing services, such as ours to efficiently print your model.
- SLS (Laser Sintering):This technology does not require support structures, therefore, it is great for very complex structures. The material is PA12 (‘Nylon’) which is flexible when thin and rigid when thick. Even though this material is relatively strong, models might still collapse if the structure is too thin. This technology requires sophisticated machines, so this type of machine is more common in industry and 3D printing services like ours than at academic or research institutes
- Colorjet: As the name suggests, this technology allows full-color prints. The material is a hardened plaster. Technically, it does not require support structures while printing, but since the material is brittle, extra support structures within the molecule can be necessary. This depends on the geometry of the model. We’re happy to make suggestions, or even do it for you if you like. As an extra service we can coat the model with an additional layer of protective clear plastic.
We hope you have fun preparing your models! Here are additional resources on how to create such models. If you know other good ones, let us know or write a comment.
- Denis Hudrisier on YouTube (using Chimera)
- Jessica Pola on ascb.org (using PyMol)
- Scott C. Meyer in the Journal of Chemical Education
- Theabion on Instructables.com
- Kalani Kirk Hausman and Richard Horne in 3D Printing for Dummies
Acknowledgment: We thank SM for providing us with the detailed Chimera instructions for this article, and we thank the University of Konstanz and the University of Tübingen for allowing us using the pictures.
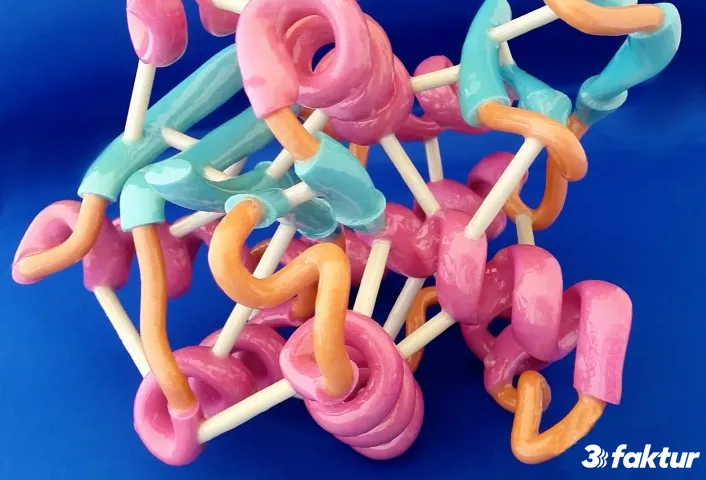
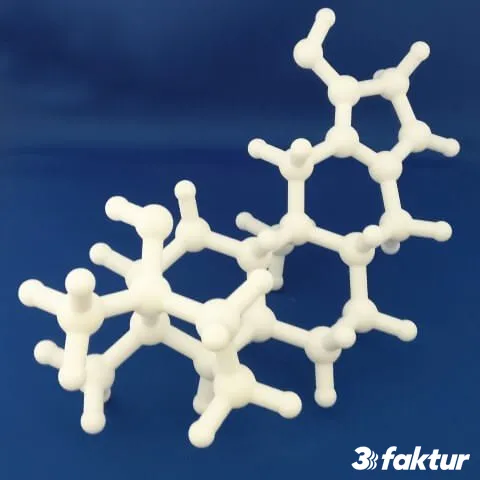



About 3Faktur: 3Faktur specializes in 3D printing, rapid prototyping, and rapid manufacturing. We utilize HP’s Multi Jet Fusion technology and offer various materials for prototyping and series production. If you have any questions about your project, feel free to contact us.
